Medics at the St. Louis campus of Washington University in St. Louis have identified transposable elements (TEs), brief DNA segments that can move around the genome, as a potential new target for cancer immunotherapy.
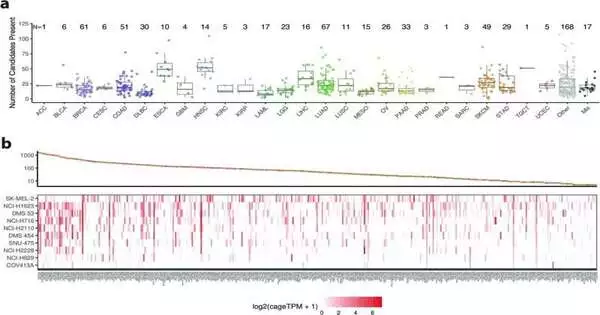
The Cancer Genome Atlas (TCGA), a database with more than 20,000 cancer samples representing 33 different cancer types and 675 cancer cell lines, was used by researchers in the study “Pan-cancer analysis identifies tumor-specific antigens derived from transposable elements” published in Nature Genetics.
The paper is concerned with transposable elements (TEs), which are present in the human genome but frequently lay dormant. These DNA fragments can insert themselves into other genes, as their name suggests. The classic illustration is a retrovirus that integrates a portion of its genetic code into the host’s genome. This kind of intrusion is anticipated by the genome, and subsequent replication and insertion are typically “turned off.”
“Drive the production of aberrant transcripts, which can be both highly expressed and extensively shared among tumor samples at times.”
According to the study,
Since cancer frequently arises from cells with damaged DNA to begin with, it frequently does not have many of the normal genetic safeguards in place when it develops. The study found that cancer reproduction enables these dormant TEs to reactivate their transposable properties within cancer cells and “drive expression of atypical transcripts that, at times, can be both highly expressed and widely shared across tumor samples.”. “.
Because of this, the researchers were able to identify TEs in 98 percent of tumors of all types, and they discovered that “these transcripts have the potential to produce new peptides, which are not present in benign tissues,” suggesting a novel strategy for attacking tumors in all cancers.
The research found 1,068 tumor-specific TE candidates that could produce TS-TEAs, or tumor-specific TE-chimeric antigens. More investigation is required to confirm that this pathway can be exploited to attack tumor cells with these TS-TEAs because there are currently cancer treatments that target particular antigen types.
Small, iterative steps of discovery are how medical science always works, and they often lead to potential cure pathways that are replete with caveats. The distinction in this case is that numerous cure-related paths have already been traveled.
The mechanism within tumors that is producing potential targets for current therapeutic strategies is what the present study has uncovered. There is a strong possibility that, with more research, a single treatment approach could be developed to address various cancer types throughout the body. This discovery might result in new therapeutic approaches for cancer treatment.
More information: Nakul M. Shah et al, Pan-cancer analysis identifies tumor-specific antigens derived from transposable elements, Nature Genetics (2023). DOI: 10.1038/s41588-023-01349-3