As they mutate, bacteria can become resistant to drugs that would kill or slow their growth by naturally adapting to various environmental stimuli.
Dr. Salvador Almagro-Moreno, a microbiologist at the UCF College of Medicine, uncovered the evolutionary origins of antimicrobial resistance (AMR) in bacteria in a recent article that was published in PLoS Genetics. His investigations on the bacterium that causes cholera, Vibrio cholerae, give insight into interpreting what conditions should happen for irresistible specialists to become safe.
“How AMR happens in bacterial populations and the pathways prompting these new characteristics are still inadequately perceived,” he said. “Antimicrobial resistance is increasing, which poses a significant threat to public health.”
“How AMR occurs in bacterial populations and the pathways leading to these new traits are still poorly understood. Because antibiotic resistance is on the rise, this poses a serious public health risk.”
Microbiologist Dr. Salvador Almagro-Moreno
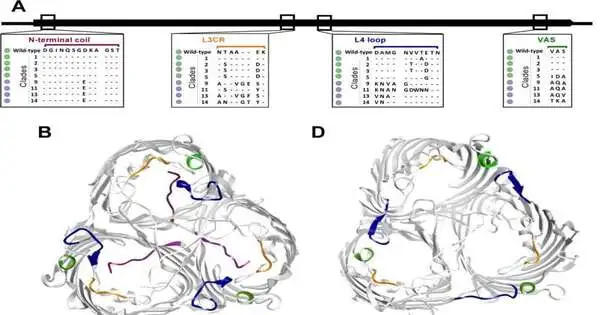
Using computational and molecular methods, Dr. Almagro-Moreno’s team discovered that several OmpU mutations in the cholera bacteria led to resistance to numerous antimicrobial agents. Dr. Almagro-Moreno studied genetic variants of an OmpU protein found in bacterial membranes.
This obstruction reminded me of antimicrobial peptides that go about as guards for the human stomach. The specialists found that other OmpU variations didn’t have these properties, making the protein an optimal framework for interpreting the particular cycles that happen to make a few microbes impervious to antimicrobials.
The researchers were able to identify specific OmpU components associated with the emergence of antibiotic resistance by comparing resistant and antibiotic-sensitive variants. In addition, they discovered that the genetic material encoding these variants as well as the characteristics associated with them can be transmitted between bacterial cells, posing a greater threat to the spread of AMR in populations under pressure to use antibiotics.
By understanding how changes happen, scientists can more readily comprehend and foster therapeutics to battle safe diseases. Additionally, environmental factors like ocean warming and pollution are being investigated by Dr. Almagro-Moreno as potential causes of resistant bacteria. “To develop a novel method for comprehending how antimicrobial resistance develops, we are studying the genetic diversity of environmental populations, including isolates from coastal Florida,” he explained.
Understanding the bacteria that cause cholera, an acute diarrheal illness linked to food and water contaminated with the bacteria, has global implications. Up to 4 million people worldwide contract the disease, and severe cases can result in immediate death.
More information: Trudy-Ann Grant et al, Allelic diversity uncovers protein domains contributing to the emergence of antimicrobial resistance, PLOS Genetics (2023). DOI: 10.1371/journal.pgen.1010490