Being as close to the source material as possible is ideal when studying the microbiology of any object, be it a molecule or a dolphin. When you’re looking into the Rube Goldberg environment of a cell’s nucleus, that can be especially difficult.
However, Princeton chemists from the Muir Lab and the MacMillan Lab used two renowned technologies in research that was published this week in Nature to direct light in the precise direction they desired.
In the process, they discovered crucial, unanticipated changes in interactions surrounding chromatin, a DNA-protein complex that functions as a sort of architecture that permits the compaction of DNA in the presence of genetic mutations that are frequently linked to cancer.
Every biological function is governed by interactions between biomolecules. A deeper understanding of cell fate results from mapping their activity. In order to maximize the benefits of in-nucleo protein trans-splicing, a protein engineering technique introduced in 2016 and since improved by the Muir Lab, researchers combined the strengths of Map and the proximity-labeling system developed by the MacMillan Lab and introduced three years ago.
“Since Map provides such very accurate information, the entire aim of Map is to comprehend biology, broadly defined, in ways you couldn’t previously. This study is an example of doing just that.”
David MacMillan, the James S. McDonnell Distinguished University Professor of Chemistry.
“The entire purpose of Map is to help you understand biology, broadly defined, in ways you couldn’t before because Map provides you with such incredibly accurate information.” The James S. McDonnell Distinguished University Professor of Chemistry and winner of the 2021 Nobel Prize in Chemistry, David MacMillan, said that this research is an illustration of doing just that.
According to this research, the biology is changing as a result of these mutations. “Before, we were driving in the dark because it was impossible to see that. It adds another brick to this wall’s construction and further demonstrates what Map will be able to accomplish. This is a true collaboration, but it’s only the beginning.”.
The combination of these technologies allowed researchers to tether an iridium photocatalyst to a protein of interest in order to study these minute interactions and how they change in the presence of mutations—all without affecting the complex microenvironment within the cell nucleus.
Researchers were able to look into this microenvironment with a level of specificity previously unattainable because the photocatalyst highlighted a radius of focus only one nucleosome wide.
Ciaran Seath, a former postdoc in the MacMillan Lab and co-lead author on the paper with Antony Burton, a former postdoc in the Muir Lab, said, “So many things in biology and disease come down to how chromatin moves and changes.”. “Viruses, aging, cancers—all the things we looked at alter how chromatin can move and respond. We reasoned that if you could watch that, you could learn about each of these various issues in a modular fashion.
“There are times when you cannot see or predict all the additional events that could possibly take place within the nucleus’ machine. We can measure these unanticipated effects now that we have paired these tools. ” Like touching a point on a spider’s web,” he said, is how it feels. “Everything is moving, as you can see.”.
The ability to install a small catalyst onto an interest protein and then map what’s nearby is what the synergy of these technologies gets you, according to Burton, in a way that causes the least amount of disruption. We were able to clarify protein interactions and intricate downstream effects at a level of detail that is very challenging to achieve with other approaches.
Most importantly, we can map how these change in response to drug treatment or mutation, opening up possibilities for academic and commercial use.
Burton, a senior scientist in chemical biology at AstraZeneca in Boston, Massachusetts, senior scientists Thomas Muir and Van Zandt Williams Jr., and Seath, an assistant professor of chemistry at the Herbert Wertheim UF Scripps Institute for Biomedical Innovation and Technology in Jupiter, Florida, contributed to the study. Professor from the class of 1965 and MacMillan
Advancing epigenetics as a field
For epigenetics, the area of biology that studies changes in gene expression, the findings have significant ramifications. Histone proteins, which protect DNA and limit access to the genome and thus play a crucial role in controlling transcription, are at the center of epigenetics.
These histone proteins have undergone mutations recently, and these mutations have been linked to a variety of cancers.
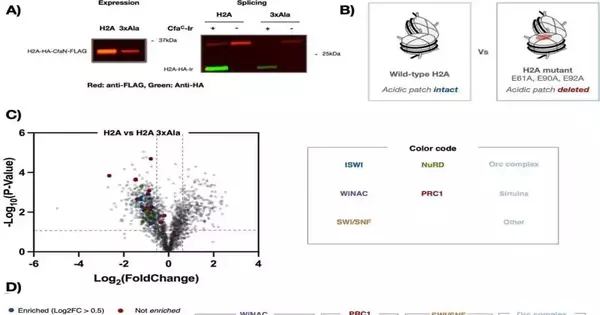
One thing Muir and colleagues examined was the addition of a histone mutation that has been associated with cancer. “What we wanted to know is what happens when that mutation is present: what can no longer be recruited, what is no longer in the area, and what new things are brought in that shouldn’t normally be there?
With the help of these technologies, he continued, “We were able to find all kinds of things related to gene regulation that change when we put this mutation on the chromatin.”. “We were able to deduce mechanistic insights about how genes are misregulated by mutations.”.
The team’s initial hypothesis when they started their research three years ago was that the cancer-related mutation causes some sort of functional loss. Biomolecules locate chromatin and bind with it to leave a transcriptional imprint. According to researchers, the mutation partially prevented that action, which resulted in cell dysfunction.
With the help of this study, which produced molecular details on how a small change in the genome can have significant effects, their hypothesis was confirmed.
More information: Ciaran P. Seath et al, Tracking chromatin state changes using nanoscale photo-proximity labelling, Nature (2023). DOI: 10.1038/s41586-023-05914-y