Global healthcare is facing a significant challenge in the form of antibiotic resistance. A new approach to tracking the evolution of antimicrobial resistance genes across bacterial populations over time is presented in an article published in the journal Antibiotics. The rapidly expanding availability of bacterial genetic sequences in public databases like GenBank is the foundation of the new computational strategy.
Ivan Erill, senior author of the study and professor of biological sciences at UMBC, says, “Our idea is that this could be used as a monitoring system.” It’s great for research that aims to learn more about what happens in bacterial genomes.”
“Our hypothesis is that this may be utilized as a monitoring system, It’s fantastic for research into what’s going on in bacterial genomes.”
Ivan Erill, professor of biological sciences at UMBC and the study’s senior author.
It only takes about an hour to analyze the sequences of all known bacterial plasmids using the code that Erill and his colleagues at the Universitat Autnoma de Barcelona, Miquel Sánchez-Osuna and Jordi Barbé, developed. Plasmids are little circular pieces of DNA that can exchange genes between bacteria. The results reveal the most prevalent resistance genes as well as their likely origin.
This kind of computational analysis is much faster and less expensive than complex systems that require global coordination among clinicians. As a result, it could be done more often to help researchers and doctors stay up-to-date on changing threats from resistance.
Erill says, “There’s going to be more and more data that you can mine this way,” pointing out that the amount of genetic sequence data that is available doubles roughly every two years. “I love it because it’s simple,” he continues. You can quickly deploy it because it’s quick.”
Genetic detective work
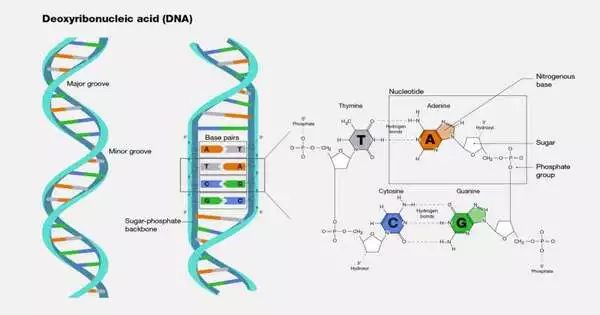
So, how does this brand-new method work? There are four bases in all DNA, including microbial DNA: A, T, G, and C. A and T pair, and G and C pair. However, the ratio of the bases varies significantly between microbial species. While some bacterial genomes may contain between 30% and 70% GC pairs, others are split 50/50 between AT and GC pairs. Erill and colleagues used this variability in a previous study to investigate the emergence of resistance to an early class of antibiotics called sulfonamides.
Through the use of plasmids, resistance genes travel from one species to another, largely maintaining their original ratio. Therefore, if the resistance gene’s GC ratio differs from that of the rest of the bacterium’s genome, this indicates that the resistance gene originated elsewhere. Because of its simplicity, this method can track the movement of resistance genes faster than clinical methods and faster than other computational methods.
It may take millions of years for the genetic sequence of a resistance gene to begin to approach the GC content of its new host if it has been present in a species for a sufficient amount of time. Erill states, “It’s basically a snapshot for what we’re looking at, which is gene movement over the last 60 to 100 years.”
Specialists spread faster
The study’s authors confirmed that resistance genes on conjugative plasmids, a type of plasmid that can easily transfer between bacterial cells, are most likely to spread using the new monitoring method. Most researchers already knew this, but proving it with the new method helped show that the method works.
In addition, the new study found that antibiotic-specific resistance genes spread the most. Since humans began using antibiotics, these genes are unlikely to have emerged naturally in any given bacterium because they typically require a large number of mutations to evolve. However, they will quickly spread if they are present anywhere in the bacterial population when the appropriate antibiotic is administered.
Erill asserts, “As soon as there is selective pressure from that antibiotic, there is selective pressure to move this thing around, because it is a bacterium’s silver bullet against that antibiotic.” This is because the antibiotic acts as a “selective pressure.” Erill explains that generic resistance, on the other hand, is less likely to spread rapidly because it only requires a few gene mutations. “Because the bacterium has probably already discovered it by the time it arrives, there isn’t a lot of selective pressure to pass it along,” he claims.
Hospitals aren’t likely the culprit
The new study also found that genes for resistance to antibiotics that are used in livestock or prescribed outside of hospitals are likely to spread throughout the global bacterial population. Hospitals are not likely to be the cause. “Resistance to antibiotics that were only used in limited settings barely grew.” “That indicates that there is less selective pressure if you use things cautiously,” Erill states.
Erill’s team discovered that most resistance genes originated from a single source and spread rather than evolving independently multiple times, which may be the most significant finding for antibiotic policy in the future. Erill says, “Resistance is in the environment, and it needs a vehicle to enter the mainstream.”
Erill argues that resistance would be significantly less prevalent if antibiotics were only utilized in hospitals as opposed to livestock and other environments. This is on the grounds that obstruction from emergency clinic utilize alone “would assume that you have normally safe microbes living in the emergency clinic as of now, prepared to pass on their qualities” he proposes. According to Erill, hospitals do indeed contain infectious microbes, but “most of the microbial diversity is in the soil and the water.” Resistance will not spread if antibiotics are unable to reach the resistant cells.
Erill states that although the new study is “more of a methods paper than a results paper,” “we believe it’s an important contribution.” It proposes a method for continuously monitoring changes in bacterial genomes over time, which may have an impact on the development of new antibiotics or treatment regimens in the future. It might even encourage restrictions on the use of antibiotics in agriculture and other settings where they can enter the environment.
Erill explains, “The best part is that other research teams can use the new method to find answers to their own questions.” You can poke at anything you want to with it and a very fine comb.”
More information: Miquel Sánchez-Osuna et al, Systematic In Silico Assessment of Antimicrobial Resistance Dissemination across the Global Plasmidome, Antibiotics (2023). DOI: 10.3390/antibiotics12020281