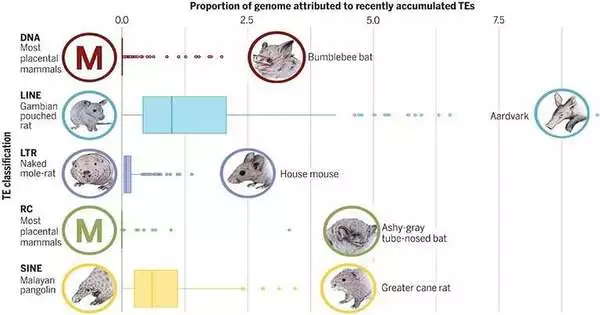
Transposable elements (TEs)—genes that can change their position within a genome, creating or reversing mutations and thus altering a cell’s genetic identity—have been identified as a crucial area of study to help uncover the evolutionary process of mammals and to better understand biodiversity by comparing the genomes of 240 living species of mammals. Liliana M. Dávalos of Stony Brook University works with the project to analyze TEs.
The findings are highlighted in two brand-new papers, one of which was published in the most recent issue of Science and the other in Molecular Biology and Evolution.
Mammals have adapted to virtually every environment on Earth over the past 100 million years. By completing comparative genomic DNA sequences from the 240 species, the Zoonomia Project, to which Dávalos is a scientific contributor, has cataloged the diversity in mammalian genomes. The group, which comprises in excess of 150 researchers around the world, distributed their long-term near-genome examination in the Science paper.
“Determining how many transposable elements of each type exist in each species is critical for understanding how transposons contribute to biodiversity. It appears simple to relate these statistics to the number of species or their ecology, but this is deceptive.”
Dávalos, Professor of Conservation Biology in the Department of Ecology and Evolution.
Dávalos investigates the biological processes that fuel biodiversity and how it changes over time. To qualitatively examine the dynamics of TEs, she collaborated with Texas Tech University’s David Ray and his lab.
The TE repertoires of 248 placental mammals are discussed in this paper. Despite the fact that TEs make up a sizable portion of all mammalian genomes, there is a great deal of variation among species. The logical group points out that relating TEs to biodiversity is not even close to basic. Moreover, with the capacity to move all through the genome, TEs can either add to biodiversity or frustrate it.
“The key to understanding how transposons contribute to biodiversity is determining the number of transposable elements of each kind found in each species.” That is misleading,” says Dávalos, a co-author of the paper and Professor of Conservation Biology in the Department of Ecology and Evolution. “It seems simple to relate these counts to the number of species or their ecology.”
“A few animal groups, like bats and whales, are, in all honesty, more firmly connected with one another than with other people, like bats and primates, so we should factor this into our measurements inside the near genomic mammalian examinations.”
More than 25,000 TE sequences were found in the mammalian set, with some mammals having a large number of TEs in their genomes, as calculated for each species over time. The normal was roughly 45%.
In general, that’s what they presumed: “Taking into account the boundless impacts that TEs force on genomic engineering, this information is a significant asset for future investigations into mammalian genomics and advancement, and we propose roads for proceeding with the investigation of these significant yet understudied genomic occupants.”
The authors of the Molecular Biology and Evolution paper used novel statistical methods developed by Dávalos to determine bat genome sequences and explain where bats stand in relation to TEs in mammals.
The research team discovered that bats are unique in that they have more events involving TE transfers from one species to another, according to Nicole Paulat, the lead author and graduate student in the Ray Lab. One component that might make sense of such abundance moves is through infections, a significant finding on how a few bat animal categories have been found to have different and sometimes risky infections.
In view of the work from the Zoonomia Venture, the two papers delineate that TEs are exceptionally dynamic across the genome of most vertebrate species, and along these lines, future examinations fixating on TEs might assist with giving solutions to mammalian biodiversity around the world. These studies may also shed light on how and why TEs disrupt mammalian genomes by altering DNA and influencing evolutionary processes and/or disease development.
More information: Austin B. Osmanski et al, Insights into mammalian TE diversity through the curation of 248 genome assemblies, Science (2023). DOI: 10.1126/science.abn1430
Nicole S Paulat et al, Chiropterans are a hotspot for horizontal transfer of DNA transposons in Mammalia, Molecular Biology and Evolution (2023). DOI: 10.1093/molbev/msad092